As the number of cases of Legionnaires' disease continues to grow, testing for this deadly bacteria became an important topic in the water management industry around the world.
Not only does routine Legionella testing ensures that water quality and safety is maintained, it also reduces the potential health risks presented by negligence, the unplanned absence of water management staff, failure of biocide dosing systems and a host of other unforeseeable events that inevitably occur within organizations.
But what testing methods are available on the market and what are the differences between them? Should only one testing method be used within the organisation, or would using various testing methods allow the business to cover more areas of risk?
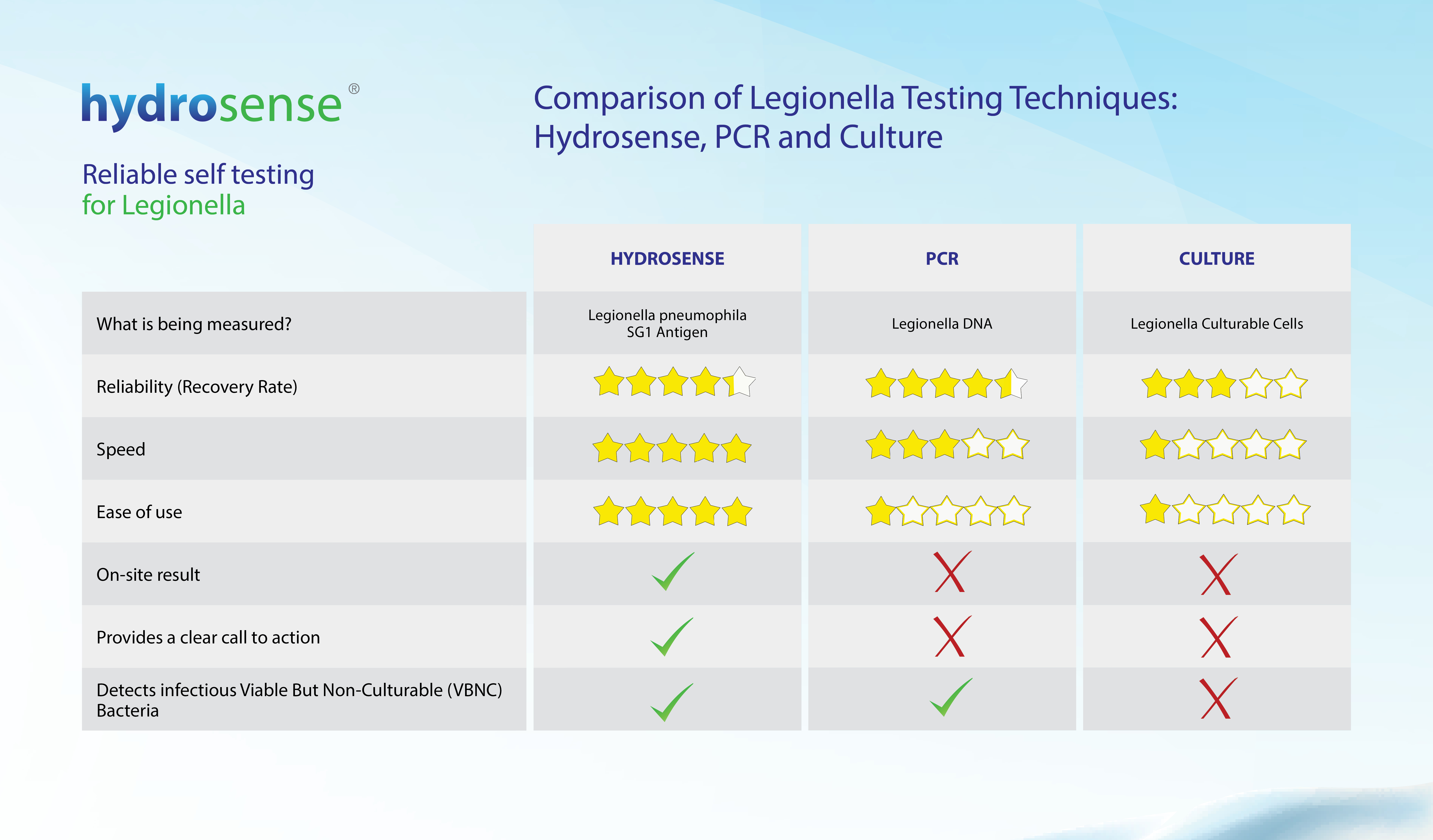
We designed this comparison table of Legionella testing methods to help you better understand differences between the antigen (Hydrosense) test, qPCR technology and the culture method.
What is being measured
Hydrosense Test: The Hydrosense test uses Lateral Flow Immunochromatographic Assay (LFICA) technology. The test is designed to detect Legionella antigen, using antibodies tagged with red nanoparticles, which bind to any Legionella pneumophila serogroup 1 bacteria found in a sample and make them visible as a line on the device. In simple terms, the test works just like a pregnancy test, except it detects Legionella antigen rather than pregnancy markers.
PCR test: PCR (Polymerase Chain Reaction) testing is a molecular biology technique in which the DNA of a microorganism is extracted and then amplified. This enables the laboratory to determine the presence and quantity of that organism’s DNA in a water sample.
Lab Culture Test: The lab culture method isolates and quantifies culturable cells of Legionella.
Recovery Rate
Hydrosense Test: The antigen test was independently validated in a paper presented at the Annual International Water Conference in 2008. The study confirmed that the recovery rate of the rapid antigen test was 80% whereas the culture method only achieved a recovery rate of 55% during the same study.1
PCR test: PCR can achieve a recovery rate of 90%2. However, this high sensitivity has to be calibrated to reflect the fact that Legionella bacteria are naturally occurring and are present in all water systems at some level. The PCR method thus requires careful interpretation to ensure the relative risk of Legionellosis presented by the system is fully understood.
Lab Culture Test: The ISO standard states that in the inter-laboratory trial undertaken, as detailed in the latest version of the standard, the Recovery Rate was only >64%3. That means there is a possibility for up to 36% inaccuracy in any culture test, even when following the ISO standard in an ISO accredited laboratory. Non-ISO laboratories are likely to have even lower recovery rates.
Speed
Hydrosense Test: The biggest advantage of the rapid antigen test is time to result. Antigen testing is the fastest method currently available in the world, for the detection of Legionella in water samples. Results are provided in 25 minutes, compared to 7-14 days in the case of lab culture, or ~24 hours in the case of PCR.
PCR test: The PCR method is a useful tool for establishing risk under emergency or outbreak conditions because it can produce a positive or negative result in hours rather than days 4, as long as an appropriate lab is close and accessible. It is more typical however, for PCR tests to take between 24 - 48 hours, particularly when shipping to the lab is required.
Lab Culture Test: Obtaining a result via the culture method typically takes between 7 to 14 days.Studies have shown that Legionella pneumophila can proliferate very rapidly, potentially doubling in population within a mere 24 hours 5. Consequently, any results received from a lab culture test could be a positive ‘call to action’ that is a week too late, or a negative result which may give a false sense of security for the system being tested.
Ease of use
Hydrosense Test: The antigen test requires no training or experience and can be carried out by anyone, anywhere. The ease and simplicity of the test can eliminate the costs associated with sending samples to the lab and is an incredibly convenient way to test for Legionella in remote areas or out at sea. Moreover, the ease of use of the antigen test enables simple, periodic sampling of water systems for the reduction of risk and is a useful addition to any water management program.
PCR test: Because the test is carried out in the laboratory, where several steps must be completed, such as concentrating viable cells through membrane filtration; sonication and heating of the concentrated cells to lyse the cells and free DNA; and purification of the DNA for the Polymerase Chain Reaction, the test is very labour intensive and complicated. It requires specialised equipment and must be conducted by trained and experienced professionals.
Lab Culture Test: Standard culture-based test can be run in any laboratory, as long as experienced laboratory scientists with the right expertise are available. The steps are complicated and require precise timing and accurate temperature control for heat pre-treatment and if an acid pre-treatment is performed, then the acid must also be prepared correctly. Because of the complexity of the lab culture methods, the difficulty in achieving reproducible and consistent results is substantial.Finally, the subjectivity in interpreting the plates leads to a lot of inter-lab variabilities.6
On-site result
Hydrosense Test: The Hydrosense Antigen test can be performed on-site, which allows detection of the bacteria in its natural environment. Field analyses also remove the risks associated with the transportation of water samples. Currently, the antigen test is the only method for Legionella detection that can be carried out on-site.
PCR test: A sample typically has to travel to the laboratory for testing, often via a 3rd party courier. In many cases, with courier-shipped samples, it’s very difficult to guarantee that best practice has been followed and that the sample hasn’t been altered
En-route the accuracy of the test can be compromised due to excessive heat, radiation or other bacteria in the sample that may dominate the Legionella present. 9
Lab Culture Test: As with the PCR method, lab test is not carried out on-site and bacteria damage can occur during the transportation process, leading to a less reliable result.
call to action
Hydrosense Test: The test is specific to Legionella serogroup 1, the strain of Legionella that is responsible for 70 to 92% of laboratory-detected cases of Legionellosis, in the United States and Europe 7. Although the test is able to detect the antigen of non-viable cells in most cases this is unlikely as antigen tends to be removed from a system quickly or will be below detectable levels when there is no active Legionella growth occurring. If the test returns a positive result then action should be taken as the concentration of Legionella bacteria is very likely to be high enough to pose a significant risk to human health. Therefore, a positive result with the antigen methods is a clear call to action for the strain of Legionella which has caused almost all known cases of Legionnaires’ disease.
PCR test: PCR provides results in a number of genomic units (GU) per litre, but an equivalence to Colony Forming Units (CFU) count (used worldwide to establish call to action) has not been established. 8Moreover, PCR enumerates DNA of both live and dead cells at very low concentrations, potentially leading to an overestimation of the actual health risk. 9
Therefore, a positive result might be hard to interpret, and risk management protocols still need to be established around the technique or the results will be unable to support the decision-making process.
Lab Culture Test: A positive Lab Culture result is a positive result two weeks too late – by the time the result is received people have been at risk of infection for two weeks. Conversely, a negative Lab Culture result provides a false sense of security as Legionella populations can double in 24 hours or less 5. A result from two weeks ago is a view of the past which holds little, if any, meaning for today.
Detects Viable but Non-culturable bacteria
Hydrosense Test: Legionella, when shocked due temperature, biocide, lack of nutrients or other stress,“shut down” functions that are not crucial to their existence, focus merely on survival and enter Viable but Non-Culturable (VBNC) state. Viable but Non-Culturable bacteria are still very dangerous and can cause an infection and lead to outbreaks of Legionnaires’ disease as they may be resuscitated back to culturable cells under suitable stimuli, such as contact with amoebae11. The antigen test can detect Legionella bacteria in this dangerous and infectious state.
PCR test: Similarly, to the antigen test, because the PCR test detects bacteria’s DNA the VBNC bacteria can be detected by this method.
Lab Culture Test: The lab culture method cannot detect Legionella bacteria which are in this non-culturable state. However, the study has shown that high percentages of the Legionella populations in water systems are not culturable. 12VBNC cells are characterized by a loss of culturability on routine agar, which impairs their detection by conventional plate count techniques. This is a major limitation of the laboratory culture method as the underestimation of total viable cells in environmental or clinical samples poses a true public health risk and inefficacy of Legionella control and management. 13
Read More About The Hydrosense Rapid Legionella Testing Kits
References:
- Polwart N., Grant R., Barnes H., Holmes E., Lindley T., Cooper A. ( 2008). 69th, Annual international water conference; 2008; San Antonio, TX, Proceeding of International Water Conference, 1, 55-68. Available at: http://bit.ly/2kDSYpb [Accessed 22 Jun. 2018].
- DiÃÅaz-Flores AÃÅ, Montero JC, Castro FJ, et al.(2015). BMC Microbiology. 15(91) http://bit.ly/2kTpQdB [Accessed 22 Jun. 2018].
- International Standards Organization. (2017). ISO-11731. Available at: http://bit.ly/2kqs97N [Accessed 22 Jun. 2018]
- Delgado-Viscogliosi, P., Solignac, L., & Delattre, J.-M. (2009). Applied and Environmental Microbiology, 75(11), 3502–3512. Available at: http://bit.ly/2lXMSjB [Accessed 22 Jun. 2018]
- Y. Buse and N.J. Ashbolt. (2011). Letters in Applied Microbiology 53, 217–224. Available at: http://bit.ly/2ksXlTW [Accessed 22 Jun. 2018].
- Infection Control (2017). Available at: http://bit.ly/2mjrEwI [Accessed 06 Mar. 2019]
- Joseph, C., Ricketts, K. and on behalf of the European Working G, C. (2010). Available at: http://bit.ly/2m0Eekg [Accessed 5 May 2018].
- Joly, P., Falconnet, P.-A., André, J., Weill, N., Reyrolle, M., Vandenesch, F., ... Jarraud, S. (2006). Applied and Environmental Microbiology, 72(4), 2801–2808. Available at: http://bit.ly/2ko06pv [Accessed 22 June 2018].
- Lee, J., Lai, S., Exner, M., Lenz, J., Gaia, V., Casati, S., Hartemann, P., Lück, C., Pangon, B., Ricci, M., Scaturro, M., Fontana, S., Sabria, M., Sánchez, I., Assaf, S. and Surman-Lee, S. (2011). Journal of Applied Microbiology 110(4), pp.1032-1044. Available at: http://bit.ly/2kQ0Kw0 [Accessed 22 Jun. 2018].
- Delgado-Viscogliosi, P., Solignac, L., & Delattre, J.-M. (2009). Applied and Environmental Microbiology, 75(11), 3502–3512. Available at: http://bit.ly/2lXMSjB [Accessed 22 Jun. 2018].
- Mendis L. Li, N., Trigui H., Oliver J.D., Faucher S.P. (2014). US National Library of Medicine National Institutes of Health. Available at: http://bit.ly/2kspNoW [Accessed 31 Jan. 2019]
- Dietersdorfer E., Kirschner A.K.T., Schrammel B., Stockinger H., Ohradanova-Repic A., Sommer R., Walochnik J., Cervero-Aragó S. (2018). Water Research 141. Available at: http://bit.ly/2kslWbs [Accessed Jan. 2019]
- Grossi M.R., Dey R., Ashbolt N. J. ( 2018). Available at: http://bit.ly/2kPTYqb [Accessed 31 Jan. 2019]